Research Program
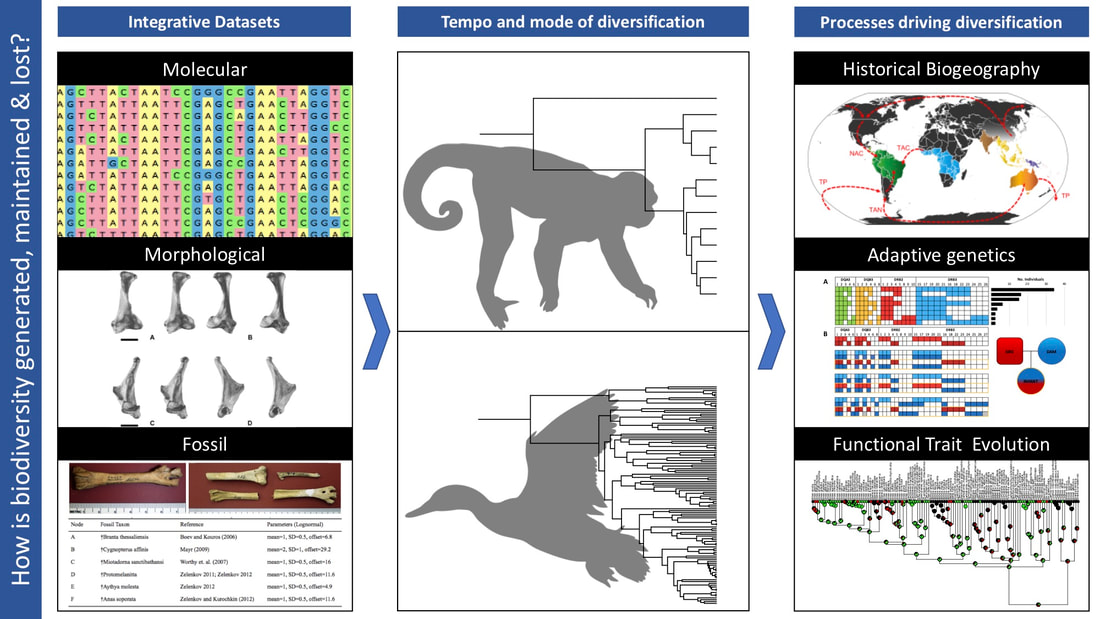
We are evolutionary biologists broadly interested in organismal biodiversity. The central goal of evolutionary biology is to understand the history of life on Earth. One critical aspect is revealing how various biological processes have interacted to shape organismal diversity through time. More specifically, how do abiotic and biotic properties of the environment and its constituent species interact to drive diversification? Key in this process is determining how organisms of interest are related. Our research program largely focuses on understanding the evolutionary history of both extant and extinct terrestrial tetrapod lineages. We integrate neontological (genomic) and paleontological (fossil) datasets to build comprehensive, robust time-calibrated phylogenies that reveal relationships as well as the tempo and mode of diversification. From there, our research continues along three main tracks:
By examining adaptive changes in genotype, phenotype and biogeography at multiple taxonomic scales within a macroevolutionary framework, we work to understand how biodiversity is generated, maintained and lost through time.
- Understanding the evolution of functional traits
- Evaluating the evolution of adaptive genetic loci in the context of evolutionary ecology
- Testing for correlations between historical biogeography and coincident physical environmental change
By examining adaptive changes in genotype, phenotype and biogeography at multiple taxonomic scales within a macroevolutionary framework, we work to understand how biodiversity is generated, maintained and lost through time.
Current Projects
Macroevolutionary Dynamics of Galloanserae
In collaboration with the OpenWings project, we are using phylogenomics to investigate the relationships among most species of the order Anseriformes (waterfowl), and a more modest sampling of their sister group Galliformes (landfowl). Together, these two orders form "Galloanserae", the outgroup to the remaining Neognathae (all living birds except the ratites and relatives). Neognathae contains the overwhelming majority of living avian diversity with more than 10,000 living species. Given the phylogenetic placement of Galloanserae and its fossil record (several of the oldest known bird fossils have been assigned to the crown and stem of this group), understanding the evolution of this clade is key to understanding early bird diversification more generally. Below are two current sub-projects related to Galloanserae macroevolution.
In collaboration with the OpenWings project, we are using phylogenomics to investigate the relationships among most species of the order Anseriformes (waterfowl), and a more modest sampling of their sister group Galliformes (landfowl). Together, these two orders form "Galloanserae", the outgroup to the remaining Neognathae (all living birds except the ratites and relatives). Neognathae contains the overwhelming majority of living avian diversity with more than 10,000 living species. Given the phylogenetic placement of Galloanserae and its fossil record (several of the oldest known bird fossils have been assigned to the crown and stem of this group), understanding the evolution of this clade is key to understanding early bird diversification more generally. Below are two current sub-projects related to Galloanserae macroevolution.
|
Project gUCE ("goose")
Anseriformes, or waterfowl, are the approximately 180 species of ducks, geese, swans and screamers. The birds occur globally, except Antarctica. We are building the most comprehensive phylogenomic tree to date for this group. One of the most interesting aspects of this work is the attempt to use ancient DNA approaches to incorporate molecular sequence data from recently extinct species in our analyses. This is part of a larger effort to examine diversification dynamics in this group by integrating molecular and morphological datasets from both living and extinct anseriform lineages (NSF DBI-1711968). With this time-calibrated phylogenetic framework complete, we are interested in looking at the evolutionary context of ecomorphologies, migration and flight, biogeography and disease ecology in this group. This work is a collaborative effort with Dr. Tracy Heath (Iowa State University), Dr. Courtney Hofman (University of Oklahoma) and Dr. Helen James (National Museum of Natural History). |
Skeletal integration and convergence in Anseriformes
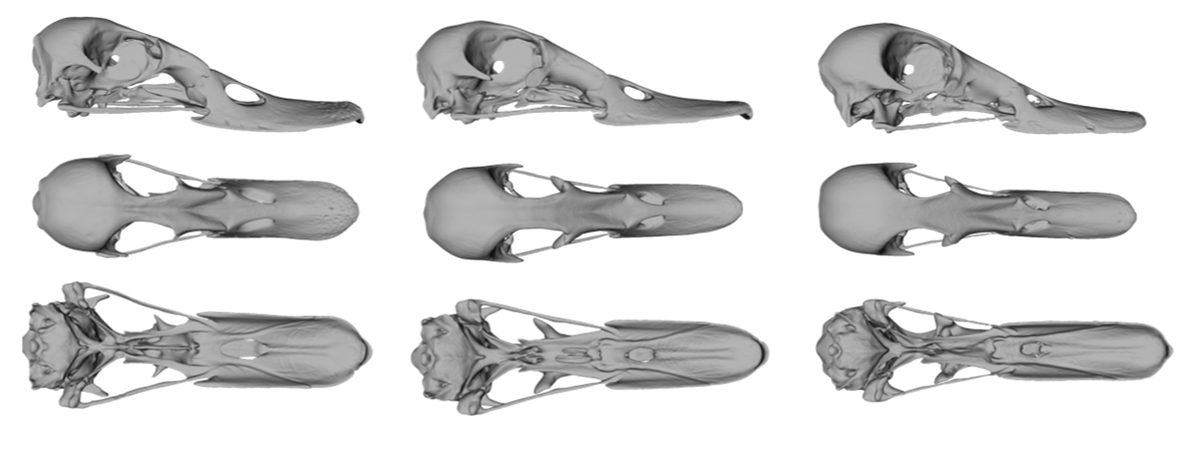
Convergence is a fascinating evolutionary phenomenon that suggests similar selection pressures in disparate groups can lead to the same or close phenotypic outcomes. Alternatively, different selection pressures can lead to the same phenotypic solutions. Studying replicate events of adaptation across taxonomic groups may reveal the selective circumstances that drive the repeated evolution of a trait or suite of traits. Our lab is interested in exploring convergent evolution in the bird order Anseriformes. Molecular phylogenetic studies of this group have revealed that previous systematic studies based on morphological data led to erroneous groupings of taxa that shared feeding modalities and related ecomorphologies. We combine phylogenomics, phylogenetic comparative methods and morphometrics to investigate the selective forces leading to convergence in the cranial and post-cranial skeleton of extant and extinct anseriform clades. We are also interested in the functional significance of particular traits, testing whether traits co-vary across the organism (e.g., modular integration) and their relative evolutionary rates through time.
Convergence is a fascinating evolutionary phenomenon that suggests similar selection pressures in disparate groups can lead to the same or close phenotypic outcomes. Alternatively, different selection pressures can lead to the same phenotypic solutions. Studying replicate events of adaptation across taxonomic groups may reveal the selective circumstances that drive the repeated evolution of a trait or suite of traits. Our lab is interested in exploring convergent evolution in the bird order Anseriformes. Molecular phylogenetic studies of this group have revealed that previous systematic studies based on morphological data led to erroneous groupings of taxa that shared feeding modalities and related ecomorphologies. We combine phylogenomics, phylogenetic comparative methods and morphometrics to investigate the selective forces leading to convergence in the cranial and post-cranial skeleton of extant and extinct anseriform clades. We are also interested in the functional significance of particular traits, testing whether traits co-vary across the organism (e.g., modular integration) and their relative evolutionary rates through time.
|
Phylogenetics and Biogeography of the Cracidae
We are also using phylogenomics to investigate the biogeographic history of the galliform family, Cracidae (chachalacas, currasows and guans). There are approximately 50 species of these birds and they occur primarily in the Neotropics. As non-migratory species, they are an interesting group for evaluating the influence of abiotic processes (climate/geology) on biological diversification in the biomes where they occur in the Atlantic Forest, Cerrado, Caatinga, Amazon basin, Andes and various regions of North and Central America. This project includes reassessments of current taxonomic delimitations in addition to analyses at phylogeographic and biogeographic scales. |
Evolutionary Genetics of the Major Histocompatibility Complex (MHC) and other Immune Genes
The MHC is a genomic region critically involved in the adaptive immune response of vertebrates. Our lab looks at the evolution and variation of MHC genes across populations, species and larger groups of organisms. The goal is to understand what selective pressures drive and maintain sequence diversity in this group of genes and how this might relate to broader diversification processes. Our published and ongoing work focuses on characterizing the diversity and structure of the MHC region across Platyrrhini, in addition to evaluating sexual selection as a mechanism for maintaining population genetic diversity in specific species. Relatedly, we want to understand how genotypes are evaluated by potential mates (e.g., via odor or sexual ornamentation). We are most interested in understanding the evolutionary context of immune genes and their function, including the possible correlations of MHC genotypes to life history traits, ecology (e.g., parasite loads) and behavior (sexual selection, sociality) in primates and other tetrapods. Below are current sub-projects related to MHC evolution.
The MHC is a genomic region critically involved in the adaptive immune response of vertebrates. Our lab looks at the evolution and variation of MHC genes across populations, species and larger groups of organisms. The goal is to understand what selective pressures drive and maintain sequence diversity in this group of genes and how this might relate to broader diversification processes. Our published and ongoing work focuses on characterizing the diversity and structure of the MHC region across Platyrrhini, in addition to evaluating sexual selection as a mechanism for maintaining population genetic diversity in specific species. Relatedly, we want to understand how genotypes are evaluated by potential mates (e.g., via odor or sexual ornamentation). We are most interested in understanding the evolutionary context of immune genes and their function, including the possible correlations of MHC genotypes to life history traits, ecology (e.g., parasite loads) and behavior (sexual selection, sociality) in primates and other tetrapods. Below are current sub-projects related to MHC evolution.
|
Chunky Monkey: Is Seasonal enlargement in male squirrel monkeys a sexually selected adaptation?
This NSF-funded (BCS-2140660) collaborative research on Saimiri collinsi (Collin's squirrel monkey) explores whether male ornamentation provides honest information to females about the genetic quality of a potential mate and thus influences her mate choice. Before and during each brief mating season, adult male squirrel monkeys gain weight producing a "fattened" appearance. Our central hypothesis is that females preferentially partner with the fattest males because such enlargement (or ornamentation) indicates that a potential mate possesses good genes to pass on to offspring. This research integrates data generated from behavioral methods in the field and molecular methods in the laboratory to examine the reproductive consequences of individual variation in male ornamentation. This work will provide new insights into the underlying molecular mechanisms of sexual selection and MHC-associated mate choice, a key area that remains relatively unexplored in primates. MHC genotyping will be performed to test the hypotheses that male fattening is a product of female choice (and thus an honest indicator of male genetic quality) or a product of male-male competition (and therefore decoupled from male MHC diversity). This work is a collaborative effort with Dr. Anita Stone (Cal Lutheran University) and Dr. Jessica Lynch (UCLA). |
Population Genetics and Phylogeography
Although most of the lab focuses on comparative research, we also have projects that address questions in single species. We are interested in understanding the drivers of population genetic structure (e.g., local adaptation, drift, isolation by distance), including in species of conservation concern. Importantly, population genetic information can provide information about the extent and distribution of genetic diversity in threatened populations and species so that conservation biologists can manage biodiversity accordingly. More broadly, populations function as the unit of evolution so genetic research at this scale provides fundamental information about evolutionary processes.
Although most of the lab focuses on comparative research, we also have projects that address questions in single species. We are interested in understanding the drivers of population genetic structure (e.g., local adaptation, drift, isolation by distance), including in species of conservation concern. Importantly, population genetic information can provide information about the extent and distribution of genetic diversity in threatened populations and species so that conservation biologists can manage biodiversity accordingly. More broadly, populations function as the unit of evolution so genetic research at this scale provides fundamental information about evolutionary processes.
|
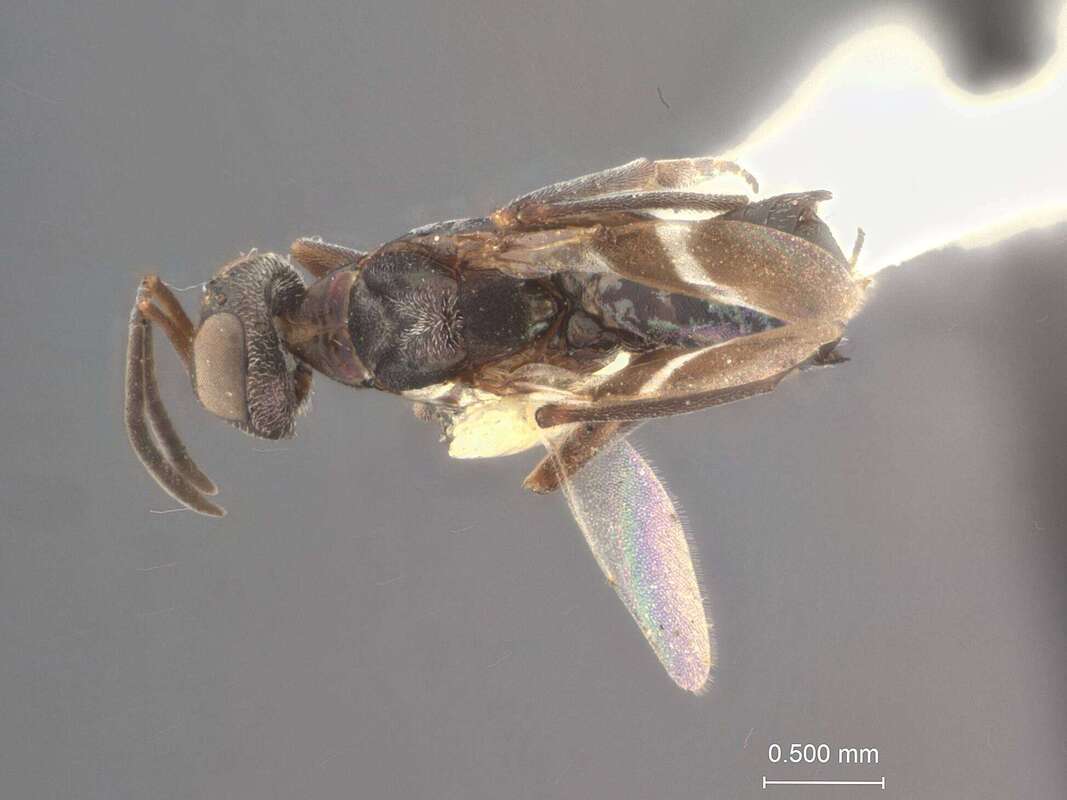
Population genetics of the parasitoid wasp, Anastatus furnissi, in the Great Lakes region.
The genus Anastatus are eupelmid wasps that are primary or hyper-parasitoids of insect and spider eggs and larvae. Some of these species are of scientific interests as biological control agents of introduced agricultural pests, like the spotted lanterfly. Anastatus furnissi is a microwasp and generalist saturniid egg parasitoid with a presumably large distribution across North America. The species was described from specimens captured in Oregon but little else is know about their biology. Interestingly, parasitism by A. furnissi has been implicated as one of the contributing factors to the decline in populations of the endangered bog buckmoth (also known as the bogbean buckmoth or Cryan's buckmoth) in upstate New York. This moth species is a unique ecological species in the Hemileuca moth radiation with limited dispersal ability. To better understanding the biology of the egg parasitoid and its contribution to native population declines, our collaborator Dr. Karen Sime has been studying the abundance and distribution of A. furnissi in the Great Lakes region and its prevalence in bogbean buckmoth populations. Our lab will be leveraging her extensive collections of these parasitoids from sites in New York and Wisconsin to understand the population genetic structure and dispersal ability of this species, as well as potential cryptic diversity across the range. |
Publications
Ghezelayagh A, Harrington RC, Burress ED, Campbell MA, Buckner JC et. al. (2022) Prolonged morphological expansion of spiny-rayed fishes following the end-Cretaceous. Nat. Ecol. Evo. http://doi.org/10.1038/s41559-022-01801-3.
Buckner, J.C., Jack, K.M., Melin, A.D., Schoof, V.A.M., Gutiérrez-Espeleta, G.A., Lima, M.G.M., Lynch, J.W. (2021) Major histocompatibility complex class II DR and DQ evolution and variation in wild capuchin monkey species (Cebinae). PLOS ONE 16(8):e0254604.
Buckner, J.C., Sanders, R.C., Faircloth, B.C., Chakrabarty, P. (2021) The critical importance of vouchers in genomics. eLife DOI:10.7554/eLife.68264.
Jones, T.L., Brenner Coltrain J., Jacobs, D.K., Porcasi, J., Brewer, S.C., Buckner, J.C., Perrine, J.D., Codding, B.F. (2021) Causes and Consequences of the late Holocene extinction of the marine flightless duck (Chendytes lawi) in the northeastern Pacific. Quaternary Science Reviews 260, 106914.
Buckner, J.C, Ellingson, R., Gold, D.A., Jones, T.L., Jacobs, D.K. (2018) Mitogenomics supports a novel taxonomic relationship for the extinct diving duck Chendytes lawi. Molecular Phylogenetics and Evolution 122, 102-109.
Lima, M.G.M, Silva-Júnior, J.S, Cerny, D, Buckner, J.C, Aleixo, A, et. al. (2018) A phylogenomic perspective on the robust capuchin monkey (Sapajus) radiation: First evidence for extensive population admixture across South America. Molecular Phylogenetics and Evolution 124, 137-150.
Lima, M.G.M, Buckner, J.C, et. al. (2017) Capuchin monkey biogeography: understanding Sapajus Pleistocene range expansion and the current sympatry between Cebus and Sapajus. Journal of Biogeography 44, 810-820. doi:10.1111/jbi.12945
Rylands, A., Heymann, E., Lynch Alfaro, J., Buckner, J., Roos, C., Matauschek, C., Boubli, J., Sampaio, R., Mittermeier, R. (2016) Taxonomic review of the New World tamarins (Primates: Callitrichidae). Zoological Journal of the Linnaean Society 177, 1003-1028.
Buckner, J. C., Alfaro, J. W. L., Rylands, A. B., & Alfaro, M. E. (2015). Biogeography of the marmosets and tamarins (Callitrichidae). Molecular phylogenetics and evolution, 82, 413-425.
Blumstein, D., Buckner, J., Shah, S., Patel, S., Alfaro, M. E., Natterson-Horowitz, B. (2015) A clinical research pathway towards developing new insights into cardiomyopathy. Evolution, Medicine, and Public Health 2015(1), 280. DOI: 10.1093/emph/eov024
Blumstein, D., Buckner, J., Shah, S., Patel, S., Alfaro, M. E., Natterson-Horowitz, B. (2015). The evolution of capture myopathy in hooved mammals: a model for human stress cardiomyopathy? Evolution, Medicine, and Public Health 2015 (1), 195-203. DOI:10.1093/emph/eov015
Buckner, J., Welsh, A.B., Sime, K.R. (2014). Evidence for population differentiation in the bog buckmoth of New York State. Northeastern Naturalist 21, 506-514.
Buckner, J.C., Jack, K.M., Melin, A.D., Schoof, V.A.M., Gutiérrez-Espeleta, G.A., Lima, M.G.M., Lynch, J.W. (2021) Major histocompatibility complex class II DR and DQ evolution and variation in wild capuchin monkey species (Cebinae). PLOS ONE 16(8):e0254604.
Buckner, J.C., Sanders, R.C., Faircloth, B.C., Chakrabarty, P. (2021) The critical importance of vouchers in genomics. eLife DOI:10.7554/eLife.68264.
Jones, T.L., Brenner Coltrain J., Jacobs, D.K., Porcasi, J., Brewer, S.C., Buckner, J.C., Perrine, J.D., Codding, B.F. (2021) Causes and Consequences of the late Holocene extinction of the marine flightless duck (Chendytes lawi) in the northeastern Pacific. Quaternary Science Reviews 260, 106914.
Buckner, J.C, Ellingson, R., Gold, D.A., Jones, T.L., Jacobs, D.K. (2018) Mitogenomics supports a novel taxonomic relationship for the extinct diving duck Chendytes lawi. Molecular Phylogenetics and Evolution 122, 102-109.
Lima, M.G.M, Silva-Júnior, J.S, Cerny, D, Buckner, J.C, Aleixo, A, et. al. (2018) A phylogenomic perspective on the robust capuchin monkey (Sapajus) radiation: First evidence for extensive population admixture across South America. Molecular Phylogenetics and Evolution 124, 137-150.
Lima, M.G.M, Buckner, J.C, et. al. (2017) Capuchin monkey biogeography: understanding Sapajus Pleistocene range expansion and the current sympatry between Cebus and Sapajus. Journal of Biogeography 44, 810-820. doi:10.1111/jbi.12945
Rylands, A., Heymann, E., Lynch Alfaro, J., Buckner, J., Roos, C., Matauschek, C., Boubli, J., Sampaio, R., Mittermeier, R. (2016) Taxonomic review of the New World tamarins (Primates: Callitrichidae). Zoological Journal of the Linnaean Society 177, 1003-1028.
Buckner, J. C., Alfaro, J. W. L., Rylands, A. B., & Alfaro, M. E. (2015). Biogeography of the marmosets and tamarins (Callitrichidae). Molecular phylogenetics and evolution, 82, 413-425.
Blumstein, D., Buckner, J., Shah, S., Patel, S., Alfaro, M. E., Natterson-Horowitz, B. (2015) A clinical research pathway towards developing new insights into cardiomyopathy. Evolution, Medicine, and Public Health 2015(1), 280. DOI: 10.1093/emph/eov024
Blumstein, D., Buckner, J., Shah, S., Patel, S., Alfaro, M. E., Natterson-Horowitz, B. (2015). The evolution of capture myopathy in hooved mammals: a model for human stress cardiomyopathy? Evolution, Medicine, and Public Health 2015 (1), 195-203. DOI:10.1093/emph/eov015
Buckner, J., Welsh, A.B., Sime, K.R. (2014). Evidence for population differentiation in the bog buckmoth of New York State. Northeastern Naturalist 21, 506-514.